|
1.
|
Introduction | ||||||
|
|||||||
|
2.
|
DemographyLab | ||||||
|
3.
|
EvolutionLab | ||||||
|
4.
|
PedigreeLab | ||||||
|
5.
|
FlyLab | ||||||
|
6.
|
TranslationLab | ||||||
|
7.
|
HemoglobinLab | ||||||
|
8.
|
MitochondrialLab | ||||||
|
9.
|
LeafLab | ||||||
|
10.
|
CardioLab | ||||||
|
11.
|
Assessment | ||||||
|
12.
|
Conclusion | ||||||
|
13.
|
References | ||||||
|
14.
|
Acknowledgements |
![]()
The Biology Labs On-Line Project: Producing Educational Simulations That
Promote Active Learning
Jeffrey
Bell, California State University
Abstract
The Biology Labs On-Line Project is an attempt to create simulations of
important biology experiments that students normally cannot perform in
typical undergraduate laboratories. All of the simulations have enough
complexity that students have the flexibility to design and interpret
their own experiments. The programs generate large amounts of data and
are fast enough for students to do multiple experiments, creating the
opportunity for students to get extensive practice at applying scientific
methods. The simulations are all written in Java and are accessed over
the World Wide Web, making them easily available to students anytime and
anywhere. Nine of the simulations, covering the topics of evolution, Mendelian
genetics, protein translation, human population demography, protein structure-function,
human genetics, mitochondrial electron transport, cardiovascular physiology
and photosynthesis are already finished, with six more planned for the
summer of 2000.
1. Introduction
1.1.
Pedagogical Motivation
A major lack in much current science
instruction at the university level is appropriate active learning experiences.
Students in many science courses still get too few opportunities to think
and reason about scientific problems. Well-designed laboratory experiments
provide the best means to give students the opportunity to learn about
a science subject while developing the thinking skills most instructors
have as a goal of their course. However, there are many limitations on
the use of real laboratory experiments in an undergraduate or high school
science course. Students in educational labs are severely limited by the
time required for most serious investigations. A typical laboratory for
a biology class will meet once or twice a week for two to three hours
each time. This time constraint is a major barrier for introductory students
as they try to learn how to be a scientist. Many important biological
experiments take weeks, months or years to carry out, putting them well
beyond the reach of the typical teaching laboratory. Many other barriers
limit the choice of experiments in teaching laboratories, including a
lack of appropriate equipment, insufficient funds for expensive reagents,
restrictions on the use of hazardous chemicals and radioactive materials,
a lack of technical skills by the students and ethical concerns.
As a result of these difficulties, most professors either concentrate on using laboratory time to provide students with an opportunity to practice being a scientist by carrying out inquiry based investigations, or demonstrate scientific principles by having students carry out carefully controlled experiments designed by the instructor. In both cases the main mechanism for students to learn the subject is the traditional lecture. A major trend in current attempts to improve science education is to try to replace static lectures with more active learning approaches. While there are many versions of "active learning," in the sciences an inquiry approach is frequently used. One version of this is the learning cycle method in which students explore a biological phenomenon before receiving any explanation from the instructor. Other aspects of the inquiry approach include having the students propose their own hypothesis and design, execute and interpret their own experiments to test their hypothesis (Lawson, et. al, 1990, Uno, 1999). The National Research Council's Science Education Standards consider this the central strategy for teaching science (NRC, 1995). In addition to learning how to think like scientists, students learn the concepts and facts of a subject better when they have to apply the knowledge. While there are many ways to make the lecture more inquiry based, (see "Handbook On Teaching Undergraduate Courses" Uno, 1999), one very useful alternative is the use of simulations (Windschitl, 1998, Uno, 1999). To address these problems the Biology Labs On-Line Project has created several Java simulations of biological phenomena that can be used to supplement traditional laboratories and lectures, providing students with many more opportunities to learn by experimentation than is possible using traditional methods.
The Biology Labs On-Line Project is a component of the California State University System (CSU) Integrated Technology Strategy (ITS), which calls for anywhere, anytime access to information. The project initially brought together biologists from throughout the CSU system and the CSU Center for Distributed Learning (CDL) to explore ways to use technology to improve learning in introductory biology courses. Later, multimedia developers from Addison Wesley Longman were added to the development team. A major goal of the collaboration was to allow students to learn as biologists do, i.e., by actively designing experiments and interpreting their results. Eliminating the time constraints of the traditional experiment, the simulations give students the opportunity to design and interpret experiments, to learn from their mistakes, and to revise and redo their experiments just like real scientists. The simulations are not designed to replace the traditional "wet labs" found in the normal biology course, but rather to extend the laboratory experience to subjects and experiments that cannot normally be done, or not done well enough, in a traditional laboratory. The simulations are also not multi-media presentations, stand-alone tutorials or on-line courses.
A key advantage of a simulation is the potential of allowing the student to design and carry out many more experiments than would be possible with real labs. This gives the student many more opportunities to practice the skills of hypothesis creation, experimental design and data analysis than can happen in the normal lab or lecture setting. A lack of underlying complexity is a problem with many of the currently available online educational simulations, which frequently allow only one real experiment. Thus, one of the design goals for the Biology Labs On-Line Project was to create simulations with enough underlying complexity that students would be able to do many different experiments. For example, the FlyLab has 29 different genes on four different chromosomes, allowing the possibility of literally millions of experiments. Similarly, the Evolution Lab, which uses two islands, each with 7 independent continuously variable parameters, provides the capability of millions of different possible experiments. In addition to the complexity in the starting parameters, each of these programs operates stochastically, so that even with identical starting conditions students will get different results. The large number of possible experiments and the speed at which each experiment can be carried out, five to ten minutes per experiment, means that students can get much more practice at designing and interpreting their own experiments than is possible in the traditional laboratory. This creates new possibilities for teaching some topics as students can now figure out the underlying principle on their own, with only minimal guidance from their instructor.
The FlyLabs introduction to the topic of sex linkage illustrates how these simulations can be used in inquiry. After doing some genetic crosses that demonstrate normal Mendelian dominance and recessive inheritance patterns, students are asked to investigate one of the X-linked traits available in the simulation. The students are not told that the trait is X-linked, nor is the concept of sex linkage discussed prior to the assignment. The students can do as many different crosses as necessary to try to figure out what is going on. The ability to carry out many different experiments is a key to making this work, as that is the only way the students can eliminate many of their initial explanations for what is happening. Even for the many students who fail to solve this problem, the experience is very helpful, making them much more attentive when the genetic explanation is given in class.
Another goal of the BLOL Project was to design simulations that could be used to learn about key concepts in biology that are not normally used in traditional laboratories because of time, expense, hazards, etc. Examples of this are the Evolution Lab and DemographyLab, which simulate processes that take place over hundreds of years; MitochondriaLab, which simulates experiments that use toxic chemicals; TranslationLab, which simulates the use of radioactive isotopes; HemoglobinLab and LeafLab, which simulate the use of complicated and expensive measuring equipment; PedigreeLab, which simulates expensive mapping experiments using dozens of human families; and CardioLab, which simulates potentially lethal experiments on human subjects.
1.2. Design
of BLOL and Comparison to Other Lab Simulations
Other laboratory simulation products
exist, comparable in some respects to the BLOL Project. Stella, for example,
is a general modeling tool, and Ecobreaker is a package that can be used
to create useful simulations of ecology processes. The Bioquest Consortium
sells a large number of simulations on CD, covering many of the same topics
covered by BLOL. These simulations differ from those in the BLOL Project,
however, in that they have certain machine requirements (Mac, PC, Unix)
and distribution difficulties. Our Java-based BLOL simulations, on the
other hand, are platform independent and not tied to use within the actual
science laboratory.
The BLOL simulations have all been created in the Java programming language, so that they can be easily accessed over the web through any standard browser. This solves the problem of widely disseminating the applications, a common problem with most educational software. The Java application provides the user interface where students set the starting parameters for their experiment and get graphical feedback on their current settings. While each of the simulations is unique, all of them share many common interface elements and functions. In some of the simulations the Java program also calculates the results, while in others the input parameters are passed back to the server, where the real calculations take place. The students receive the results through the Java application.
The downside of using Java is that only individuals and schools with fairly new computers and software (Netscape 3 or better, etc.) and an internet connection can use the software. Another disadvantage of using Java is the inability of Java programs to save to disk or print. This limitation has been overcome through the use of a notebook that can be exported to a web page. All of the data tables, such as numbers of different types of progeny, or results of statistical calculations, can be imported directly into the notebook. After typing in their comments, students can export the notebook to a web page for printing, or to email to themselves or an instructor. The web page is temporarily stored on the server. Graphical images produced by some of the programs are also exportable to the notebook, where they can then be printed.
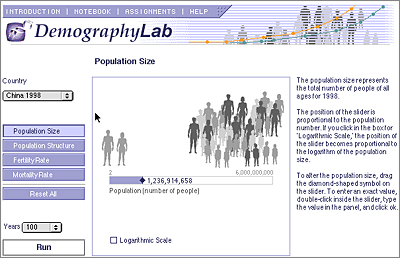
All of the programs share some common user interface elements, including a title bar with links to an introduction to the lab, help, sample assignments, the notebook, etc. While there is much diversity in how the different labs operate, most of them start in an input mode where the students design their experiment by adjusting different parameters, after which they run the simulation. The program calculates the results of the experiment, usually in a minute or less, and then presents the results in the output mode. In this mode there is a tabbed interface where the students choose which type of output they wish to view, a table of the data, a graph, the input values, etc. After analyzing their results they can import them into the notebook and then go back to the input mode to design another experiment. This ability to go back and forth quickly between the design of an experiment and the results is one of the powerful advantages of a simulation approach to teaching science.
1.3. Availability
For simulations to truly be useful as a primary
mechanism for teaching the concepts of a course, they need to cover the
majority of the key concepts in the course. The BLOL Project has produced
nine different educational simulations covering the subjects of evolution,
Mendelian genetics, protein translation, human population demography,
protein structure-function, human genetics, mitochondrial electron transport,
plant photosynthesis and respiration, and cardiovascular physiology. Current
projects in progress and due to be finished by the summer of 2000 will
simulate enzyme kinetics (EnzymeLab), the use of transgenic mice to study
developmental genetics (TransgenicLab) , phylogenetic reconstruction (CladisticsLab),
population genetics (PopulationGeneticsLab), metabolism (MetabolismLab)
and population ecology (PopulationEcologyLab). Thus, a significant portion
of the major concepts introduced in a typical introductory biology course
could be taught using a combination of real laboratories and these simulations.
![]()